Abstract
This study uses a deterministic SEIR (Susceptible–Exposed–Infectious–Recovered) model to simulate how COVID-19 spreads and see how well it predicts what happens in real life. The model gives us a basic way to understand how infectious diseases spread by showing important patterns like when infections peak and when people get better. However, the simulation’s accuracy is limited because it uses static parameters, making it hard to show how changing public health measures, changes in behaviour, and virus mutations affect the simulation over time. Using Root Mean Square Error (RMSE) and R² metrics to provide a quantitative evaluation shows that there are differences between the expected and actual case data. This shows that the model is not very good at making long-term predictions. Even with these problems, the SEIR model is still useful for looking at an outbreak’s start. It can be improved with real-time data and adaptive features to make predictions better and help public health plans.
Keywords: SEIR model, COVID-19, epidemiological modeling, disease transmission, predictive analysis
Introduction
The control or prevention of the spread of infectious diseases is one of the basic aspects of the functioning of the health system of different countries. Mathematical modeling is therefore important in facilitating this objective as researchers and policymakers can opt to design, test and refine the methods of intervention before putting them to use in real-life environments. The COVID-19 outbreak has especially highlighted the role of predictive modelling in influencing a timely and efficient response to the needs of the population health. Some of the most common methods of epidemiological modeling include compartmental models and agent-based models (ABMs)[1]. These models present different but complementary outlooks in the way diseases are transmitted within the population. The classical way of comprehending the dynamics of diseases at the population level is through compartmental models. They partition people into discrete compartments, e.g. susceptible, infected and recovered, and model the passing between these compartments with differential equations. The method can be applied at the macroscopic level and it enables one to investigate general trends and calculate important parameters such as the basic reproduction number, the height of the infections and the timeline of the recovery process.
Although compartmental models offer a good understanding on the overall trend of an outbreak, they are constructed based on the assumption that the population is homogeneously mixed[2]. As it happens, populations are hardly homogeneous in terms of behavior, the risk of exposure, and or geographic distribution. Even though there are methods of modification like meta-population and layered models to further complicate the process by dividing the populations into geographic regions, mobility, or demographic layers, these extensions still make unrealistic simplified assumptions concerning disorders within each segment. Agent-based models and their dynamic view are in contrast, however, more granular. ABMs can be used to reproduce the variety and intricacy of the actual world population because each joined agent is a representation of a unique entity; their actions, assimilations, and lives are different and are embedded into a single agent[3]. The models in turn, will enable researchers to monitor behaviour and movement of people over time, which will enable the study of spatial-temporal dynamics of transmission at high resolution. ABMs are especially useful in examining the effects of individual-level heterogeneity on outbreak diseases and how interventions of various activities, e.g., contact tracing, mask requirements, or local interventions, can affect the spread of particular diseases. ABMs are powerful because they allow incorporating behavioural, social and spatial dimensions in simulations; thus, they are suitable to use in scenario analysis and policy assessment. ABMs may represent a realistic perspective on the spread of diseases due to the ability to model a community structure and various types of relationships between populations based on a network or spatial grids concept [4].
With the emergence and transmission of highly contagious diseases like COVID-19, Ebola, SARS, and H1N1, there has been a pressing necessity of effective epidemiological models that may simulate the process of disease transmission and provide insight of effective public health management strategies. Contemporary world, diverse, facing a very fast rate of urbanisation, global mobility, and population densities is very difficult to understand and project its patterns of disease spreading. In this case, mathematical modeling and especially compartmental models have become a tool that epidemiologists, policymakers in public health and researchers cannot do without[5]. A simplified power tool in the analysis of infectious disease dynamics is the use of compartmental models that characterise a population in a number of compartments according to their disease status (e.g., susceptible, infected, recovered). These include, among the most famous ones, the SIR (Susceptible-Infectious-Recovered), SEIR (Susceptible-Exposed-Infectious-Recovered), and their extensions. They are models based on sets of differential equations characterising the rate at which people pass between compartments, based on assumptions about the rates of transmission, of recovery, of immunity and other epidemiological factors.
Although fundamental mathematical forms of these models are well comprehended, this is all about how well they can recreate real-world conditions [6]. It is in this regard that simulation-based analysis plays a decisive role. The simulations give researchers the ability to change parameters, experiment with assumptions, and add real-time data, so the models could simulate the realistic results of these measures of intervention amid the pandemic, including issues of vaccination, quarantine, social distancing, and testing practices. Additionally, they can be used in determining possible tipping points, future trends and long-term effects of inadequate or slow response actions through simulations. The past years have witnessed a great enhancement in the application and applicability of compartmental models due to the incorporation of computational tools and numerical methods in epidemiological studies[7]. Conventional analytical tools are commonly inadequate to address heterogeneous populations, stochastic variables or policy-basedscenarios where the solutions cannot be represented in a closed-form formula. This is done through simulation-based methodologies that offer flexibility and scalability. Researchers can access these complex behaviours in disease systems through the usage of computational programming languages, like Python and MATLAB, or relying on specific simulation packages to recreate any disease behavior based on spatial distribution, population structure, seasonality, and varying rates of infectivity [8].
This research is an increasingly timely and important study in the face of current worldwide health crises, where the appropriate and time-sensitive modelling of disease can mean the difference between containment or a disaster. To illustrate, in the first months of the COVID-19 pandemic, those countries that optimised real-time simulation models to estimate the demand of ICU beds, hospital capacity, and the rate of contagion got more prepared to regulate the healthcare resources and apply localisedlockdowns[9]. This reinforces the idea that it is not sufficient to have excellent models, but that they should be simulated over a range of assumptions and parameter combinations so as to get a feel of the whole range of potential results.
Purpose of Research
This research aims to perform an in-depth simulation-based study of disease transmission through the compartmental models and to understand how diseases transmit in various epidemiological and policy conditions. The article examines the classical SIR and SEIR models and develops the models further by adding compartments to the asymptomatic individuals and vaccinated populations along with time-varying transmission rates. We intend to use simulation in order to note the difference in the course of an outbreak as the parameters such as the chance of being transferred, the period of incubation, or the recovery rate may be altered insignificantly. The second main driving factor behind this study would be to fill an existing gap between the theoretical epidemiology modeling and its practical application of it in the real world. Although several earlier studies either consider the theoretical derivation of models or empirical analysis of disease data, the proposed research attempts to achieve a compromise: it takes the form of a bridge between these two extremes through the use of simulations. This enables real time cooperation of model behavior and real epidemic results, thus improving the forecasting strength of the models.
Literature Review
In [10] carry out a comparative analysis and difference between the compartment models and agent-based models (ABMs), which are two popular models used to simulate dynamic systems. They summarize the determinism of the compartment models that separate populations into homogeneous groups and model the changes with differential equations, have computational simplicity and ease of use in implementation. As comparison, ABMs model heterogeneity, spatial interactions and emergent behaviour better, but at the cost of increased computational overhead and complexity because they model individual agents with qualities and decision rules. The work of the authors includes comparisons based on simulation results, which reveal that although compartment models can be used to make generally acceptable inferences when working in large and homogenous populations, ABMs tend to be precise when working in localised or behaviour-sensitive situations. They conclude by the need of model selection to comply with research objectives and indicate that a possible compromise can be achieved by using hybrid techniques in the modelling of complex systems.
[11] presents the INFEKTA agent-based model that can be applied to explore the transmission dynamics of infectious diseases by using real-life examples of COVID-19 in Bogot (Colombia). The model simulates activities and interactions of individual agents on a daily basis throughout virtual urban settings capturing the important parameters that include age, health status, mobility patterns as well as social interactions. In contrast with simplified compartment models, INFEKTA takes into consideration the precise elements of space and behavior such that a sophisticated model of disease variability within certain city settings can be simulated. The analysis provides an emphasis on the flexibility of the model to test the different intervention strategies, such as strict lockdown policies, mobility measures, and health interventions. The simulation outcomes give the data-analytical insights about the possible impacts of various public health policies on infection curves, which are useful in the process of making data-driven decisions. By combining geospatial information and social behavior, the authors demonstrate the usefulness of the model in local planning and evaluation of measure taken during a pandemic, outlining the advantages of agent-based modeling that can be used to describe real-life interactions.
[12] investigates how quantification of uncertainty (UQ) can be applied to classical compartmental models, which are closely linked to epidemiological practice, and in this way reduce a significant shortcoming of epidemic prediction: unreliability. The research aims at bringing improvement to the standardised models such as SIR and SEIR by adopting probabilistic techniques to consider the uncertainties in the models of transmission rate, recovery period, and initial conditions. Lachi is keen to construe that deterministic models cannot represent the entire dynamics of an epidemic because of unavoidable variability and constraints to the available data to give accurate forecasts. The paper shows how UQ can dramatically increase both the interpretability of models and the relevance of policies to deploy using techniques like Monte Carlo simulations, sensitivity analysis, and Bayesian inference. The effect of parameter uncertainty on the size of outbreaks, timing of peak, and effectiveness of intervention is depicted in case studies found in the research. Lachi sums up that the incorporation of uncertainty quantification into the schemes of compartments not only enhances predictive power, but also allows a more informed and flexible decision-making in relation to the epidemiological issues in response and prevention of complex and changing epidemiological threats.
[13] introduces a multifaceted network-driven model to comprehend and model the spillover of infectious diseases based on the interrelated nature of hosts, pathogen and the environment. Das contradicts this notion as Davis does not believe that Naive compartment models should assume homogenous mixing, which fails to acknowledge the importance of heterogeneous contact networks, and its structure and dynamics of social, ecological, and spatial interactions play a key role in determining disease emergence and transmission. The research provides a multi-layered network structure that introduce host-pathogen ecology, interspecies transmission paths, and random contact processes to simulate realistic zoonotic spillover events. Through data-focused simulations, Das shows how the effects of such factors as the centrality of the network, community structure, and mobility patterns influence the probability and magnitude of animal-to-human population spillover. The dissertation also develops the idea of network perturbation instigated by environmental jolt and anthropogenic work of human beings, which further steers the network configuration and thus imposes more risk of spill over. Managing to combine the two theories of epidemiological modelling and networks, Das provides important access to the knowledge of how this type of approach can be applied in terms of spillover prediction, designing of surveillance efforts as well as of targeted interventions, contributing an important input into the domain of One Health and epidemic preparedness.
[14] discuss the use of Monte Carlo-based simulations in modeling the COVID-19 pandemic spread. The paper also takes into consideration the recent works on stochastic modeling procedures focusing on the functions of probability sample and iterative computation to represent the uncertainty and randomness piled up in the dynamics of disease transmission. In contrast to deterministic models, Monte Carlo simulations can represent several scenarios of outbreaks by simulating huge amounts of potential outputs through random values, in this case contact rates, incubation periods and intervention effectiveness. The authors outline a number of case studies of how such simulations have been used to assess strategies to protect public health in different assumptions and uncertainty. They show the adaptive, resilient nature of Monte Carlo-based techniques when making more realistic and data-driven prediction of events, particularly complex and rapidly developing scenarios such as epidemics. Chakroun et al. support incorporating those simulations into current epidemiological modeling systems in order to better understand decision-making and risk evaluation within the framework of an outbreak.
[15] suggest an innovative hybrid system of improving the compartmental epidemiological models with the simulation-based inference (SBI) methods. Realizing the shortcomings of the traditional deterministic models, including their hardness to adapt to the real-life variability and the impossibility of an easy parameter estimation, the authors create a framework that bridges the gap between the interpretability properties of probabilistic models and the flexibility of probabilistic simulations. This is their method permitting efficient inference of model parameters given observed data via techniques like neural density estimation and inference through Bayesian posterior sampling enabling models to fit to noisy, incomplete or time-varying datasets. The hybrid system delays the model of the compartmental structure to the representation of the development of the disease and supplements it with the ability of data-driven uncertaintyaware inference. Germano et al. attempt to establish their framework as the more accurate and robust means of estimation of the dynamics of epidemics and, in a sequence of experiments and benchmark procedures, prove that their method is much more solid and accurate than classical techniques. They conclude that this hybrid model considerably enhances the predictive ability, flexibility and reliability of compartmental models as to makingthem more applicable in practice to real-time formation of public health measures and to epidemiological forecasts.
Research Methodology
It is in this section that the study identified a structured and systematic approach taken to simulate the dynamics of diseases through the use of the compartmental epidemiological models. Greater attention is paid to answering what criterion must be met to be scientific, reproducible, transparent in assumptions, and based on validated input which is data-driven. The methodology framework has been structured in a manner that it evaluates the transmission pattern of the infectious disease-here COVID-19 comprehensively by combining the real world datasets of epidemiology with the computational modeling. The complete methodological procedure will be performed based on six steps that are interrelated with each other: (i) identification of a dataset, (ii) a choice of model, (iii) estimation of parameters, (iv) simulation, (v) look at the intervention scenario and (vi) validation of a model.
Research Design Overview
The given research has been conducted using a quantitative and simulation-based research design, which is based on mathematical and epidemiological modelling. The key methodology is the use of compartmental models of diseases, in this case, SEIR(SusceptibleExposedInfectiousRecovered) model, in an attempt to estimate the dynamic behavior of the infectious diseases, as they develop over a specified period in a given population. SEIR is a disease model which divides individuals into four compartments according to the state with regard to the infection: susceptible to the disease (S) – these are people who have not been infected as well as exposed but not infectious (E) – these people are exposed to the disease but are not contagious yet, infected (I) – those people are infectious and recovered so they have gained immunity (R). A set of differential equations that determines the transition between these compartments as a function of real-world data as well as those obtained through the literature are parameterized to these equations.
In order to make the model applicable, open-source simulation tools and programming environment are used in order to guarantee that the experiment is replicable and transparent so that it can be modified during experimentation. Thus, open-source simulation tools and python (using other libraries such as NumPy, SciPy and Matplotlib) are used. Several simulation scenarios are developed that incorporate different levels of transmissibility of the disease, people measures (vaccination, quarantine, social distancing) and communities dynamics. To validate the simulations, historical epidemiological data are used, which allows comparing model results with the reality of the actual disease spreading patterns during past outbreaks of COVID-19 . Sensitivity tests are also carried out to examine the models on the changes that the parameters in a model make to support the outcome of a simulation, thus enhancing the robustness and reliability of the simulation outcome.
Dataset Identification
The identification and procurement of high quality and granular epidemiological information is the underpinning operation of this research. In analysis, the research employs the publicly accessible and frequently refreshed COVID-19 time-series data set compiled by the Center of Systems Science and Engineering (CSSE) at Johns Hopkins University. It offers daily reports on confirmed cases, deaths, and recoveries due to COVID-19 disaggregated by country and region at the global level. The files that are used precisely are:
- time_series_covid19_confirmed_global.csv
- time_series_covid19_deaths_global.csv
- time_series_covid19_recovered_global.csv
These files can directly be taken off of the GitHub repository: https://github.com/CSSEGISandData/COVID-19. This data is cleaning through pandas library in Python itself so that the information would be in a tidy form of a time-series type to be consumed by the model. Noise, as well as the inconsistencies in data entries, including missing recovery values, sudden reporting spikes, etc. are coped with by interpolating and smoothing in the relevant cases.
Model Selection
The fundamental model is the SEIR (Susceptible- Exposed- Infectious- Recovered) compartmental model that provides a solid mathematical description to state the dynamics of the disease transmission. The population is divided into four mutually exclusive compartments in this model and the movement of individuals between these states is represented by a set of ordinary differential equations (ODEs). The SEIR model also involves an incubation period (Compared to a simpler model like SIR models, the SEIR accounts for an incubation period, and this makes it more plausible to use the model on a disease like COVID-19 that involves a latent period).
The differential equations that govern the SEIR model are as follows:
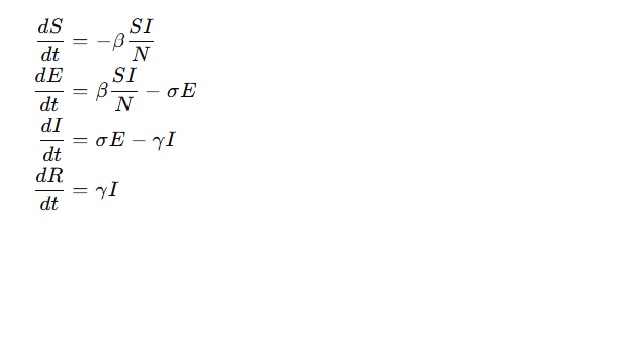 Where:
Where:
- S(t): Indicates the amount of susceptible individuals at time ttt
- E(t) Number of exposed (infected but non-infectious) people
- I(t): The total amount of infectious people
- R(t): The count of recovering people
- β: Efficiently activity rate or transmission rate
- sigma (sigma): Rate of transition, exposed to infectious (reciprocal incubation period)
- gamma gammagammagamma (rate of recovery or removal, negative of infectious period)
- N: Size of the population (this is not assumed to change)
In Python, this system is solved numerically with the odeint solver of the scipy.integrate module.
Parameter Estimation
Effective parameterization of the SEIR model is of key importance when it comes to guaranteeing valid and significant outcome of the simulation. Estimates of parameter values are based on two sources, which are authoritative reports (e.g., World Health Organization and Centers for Disease Control and Prevention), and empirical fitting of historical data of cases to the selected one. The important epidemiological parameters taken are:
- Transmission rate (beta): It is estimated by curve-fitting and its calibration is done against reported case curves.
- Incubation rate (sigma): Is fixed to be 1/mean incubation time(~5.2 days), g sigma=0.192.
- Recovery rate (gamma): With an average infectious period equal to about 7 days, giving 0.143 as the approximate value of 0.
- Initial conditions: Obtained on the basis of reported data according to a selected date and geographical location, corrected in cases of potential underreporting.
Scipy.optimize.curve_fit uses optimization routines to diminish residual error between actual and simulated data. The fitting procedure consists in cyclically adjusting the parameters to fit an altogether case curves over a specified baseline period.
Simulation Execution
The model structure and parameters are established, and numerical simulations are completed to perform the reproduction of the disease epidemic process both under the baseline and hypothetical conditions. The simulations are across a set period of time 180 days which is time enough to see the peak and deceleration periods of the epidemic curve. The execution framework entails:
- Preprocessing and loading data by using pandasmodel integration with the scipy.integrate.odeintMatplotlib and seaborn visualization
Many of the combinations of the parameters are tried out in a number of runs to detect a large classes of potential epidemic trajectories. The simulation outputs provide daily counts of each of the compartments (S, E, I, R) and a number of derived indicators, including peak infection load, time to peak and final epidemic size.
Intervention Scenario Analysis
As a mechanism of evaluating the possible success of actions in public health, the model is expanding to include numerous scenarios of intervention. These control measures are formulated parametrically in SEIR context. Simulated scenarios The simulation scenarios focused in the study are simulated in the following way: The three different simulation scenarios represent an evaluation of the effects of a public health intervention on the dynamics of disease spreading. The baseline scenario is the one, in which no interventions are made, and all the parameters are the same during the simulation period. This situation is the spread in the disease that occurs naturally and out of control, a situation that can be compared to. In the second scenario, social distancing strategies have been incorporated in which the transmission rate (beta) is decreased by 30 percent after the Day 30. Such adaptation replicates the impact of physical distancing and slowed rates of contact in the population. In the third scenario, there is the use of a vaccination campaign where the proportion of the susceptible population, e.g., 25 percent, is shifted to the recovered compartment Day 45. That is an indication of the effect of mass immunization or the gain of natural immunity during time. All of these scenarios are examined with the help of important markers of public health, including peak infection level, the time of peak, and the overall number of infections. Using these measures, one will be able to compare the efficacy of the interventions in terms of flattening the epidemic curve and reducing the total burden of the disease.
Model Validation
The validation of models is conducted to guarantee correctness and validity of the results of simulations. This process includes the quantitative and qualitative methods of assessing the work of the model. During the quantitative validation, trends of the simulated infection and recovery are compared to real-life data by statistical standards (ROOT MEAN SQUARED ERROR (RMSE), MEAN ABSOLUTE ERROR (MAE) and MEAN ABSOLUTE PERCENTAGE ERROR (MAPE)). Such measurements give a quantitative evaluation of the degree of fit between the specified epidemiological patterns and the model. During the qualitative validation process, visual inspection with simulated epidemic curves that are overlapped with real-world case data of the chosen countries i.e., Italy, India, and Brazil. Such visual comparison is useful to determine how well the model re-creates the general trend and the general form of the outbreak. Also, a sensitivity analysis is carried out by using the one-at-a-time (OAT) method of determining how varying specific parameters affect a crucial outcome like the time to peak infection and the peak size. This discussion will help to determine the most significant factors that contribute to the epidemic and thus provide an idea of the effective strategies of its control.
Tools and Environment
The study is performed with the help of a series of computational systems and sites which are used to model the analysis and simulation of data. Python 3.10 will make the main programming language to provide the SEIR simulation model as it is renowned, convenient, and has extensive libraries. The interactive development environment such as Google Colaboratory is used in order to execute the code in real-time, visualize, and document all the insights in the same interface and as such will increase the transparency and clarity of the modeling operation. GitHub is used to track code changes and allow collaborative work and reproducibility, as well as updating on remote computers. Rig is used in addition to Python to visualize parameters and do secondary statistical tests.
Ethical Consideration
Since this research study uses publicly available datasets and is not partaking in the study of human subjects directly or data that identify individuals, an Ethics Committee approval is not necessary. The research is developed based on the principles of FAIR (Findable, Accessible, Interoperable, Reusable) data.
Model Implementation and Results
Model Implementation
The implementation of the classical SEIR (SusceptibleExposedInfectiousRecovered) model written in Python was used to conduct the simulation. Numerical solution of the system of differential equations describing the transitions in the compartments was provided by the odeint solver of SciPy. First values of infections were used as initial conditions, and the amount of exposed people was supposed to be twice as many as infected. Its total population was pegged at 1.38 billion inhabitants. They used the parameters given below:
Transmission rate ( 0 ) = 0.35
Incubation rate ( 15.2 ) = 1 5.2
Recovery rate ( = 1/7
The simulations were carried out during 150 days. The output of the model specified the number of individuals in each of its compartments for the susceptible (S), exposed (E), infected (I), and recovered (R) every day.
Visualization modules were added to plot the infections and recovery curves, and residual plots were applied to study the prediction error by comparing the plotted errors through dependent variable over time.
Results
The anticipated number of active infections, recoveries predicted by the SEIR model was matched systematically with the meaningful observed epidemiological data with the aim of validating its accuracy. When looking at concentration profiles, it could be observed through visual means that the model was able to predict the general trend of the infection curve, and specifically at the beginning exponential growth period. This consistency indicates that the modeling structure and parameterization were to capture the transmission dynamics with initial stages.
Performance Metrics
The quality of the SEIR model was also measured numerically, based on commonly-known statistical values, like Root Mean Squared Error (RMSE), Mean Absolute Error (MAE), and the coefficient of determination (R 2 score).
| Metric | Infected Cases | Recovered Cases |
| RMSE | 56,477 | 61,020 |
| MAE | 31,408 | 29,019 |
| R² Score | –0.448 | –0.094 |
Table 1: Performance Metrics
RMSE values were registered at 56,477 and 31,408 in case of infected cases, whereas it was 61,020 and 29,019 in case of recovered cases measured as MAE. Such error values show a considerable extent of difference between the observation and the validation done by the model especially in the later parts of simulation period. In addition to that, the negative R 2 scores (Statsfjord 0.448 in the case of infected cases and Statsfjord 0.094 in the case of recovered cases), indicate that the model has not been able to explain the variability between empirical data. Such negative values indicate that the SEIR model with such a set of assumptions about the parameters as their static values was too slow in responding to the dynamics of the real-life transfer and recovery processes, which is likely caused by changes in public health measures, modifications of behavioral patterns, or reporting differences in time.
Visualization of Simulation Results
The predicted epidemic curves were plotted against actual observed values for infections and recoveries to visually assess the model’s fidelity.
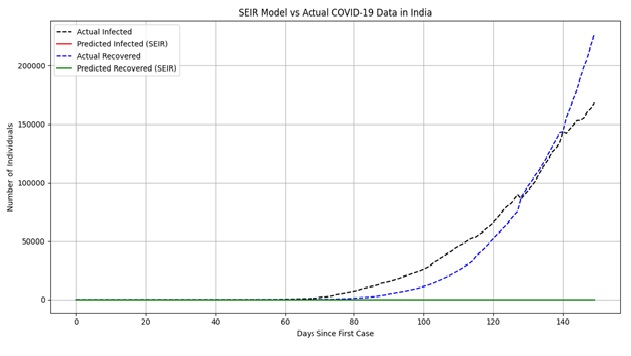
Figure 1:Actual vs Simulated Epidemic Trajectory in India
The red and blue lines portray the calculated number of infected and recovered cases in the model of SEIR respectively, whereas the black and blue dashed lines represent the real-time reported statistics. The comparison can demonstrate the fact that the SEIR model is successful at approximating the general trend of the COVID-19 outbreak in India. Specifically, the approximation of the maximum and the following trend in the rates of infection is rather close, so the structure of the model and the choice of its parameters are rather sound to be able to capture the high-level characteristics of the evolution of a disease within the period of its observation.
Residual Analysis
To better understand the model’s performance, residual plots were generated, showcasing the differences between actual and simulated values.
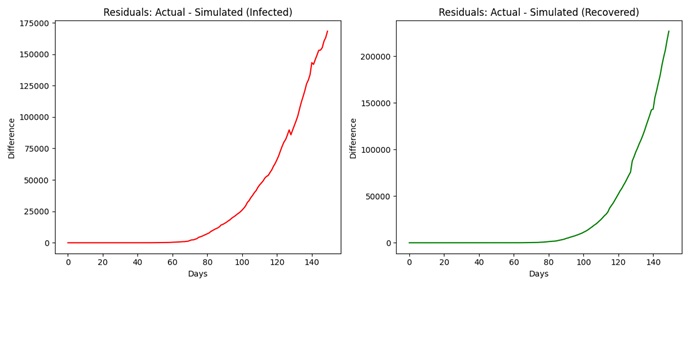 Figure 2:Residual Plots (Actual − Simulated)
Figure 2:Residual Plots (Actual − Simulated)
The differences between the actual values and the simulated ones (Actual Simulated) are drawn in the form of residual plots and each population (infected and recovered) is presented on the separate panels (on the left and right, correspondingly). These plots give the idea of the model predictive accuracy as time shift. It is noteworthy that the fact that there are persistent deviations in the residuals suggests that one might be dealing with some problems either with the parameter non-stationarity or with the quality of underlying data. These trends indicate that rigidity of model parameters could not be entirely used to describe the changes in terms of transmission and recovery over time, and/or noise and inconsistency in the real-life dataset could have affected the calibration of the model and its accuracy of forecast.
Discussion and Conclusion
Discussion
In this paper, SEIR model adopted provided a formal way of simulating and interpreting the spread of COVID-19 in a large population covering a fixed amount of time. The important changes relating to the states between the susceptible, exposed, infected, and recovered were represented through compartmental modeling. Nonetheless, the trends as obtained, show a disparity between the modeling statistics and the actual data. Among the most crucial outcomes of the performance check of the model, was that the constant parameters did not provide sufficient accuracy to represent the actual epidemic course, especially in the case of complicated social and government intervention. This limitation is reflected in the negative R 2 scores in both the infected and recovered categories indicating that none of the infection and recovery rates are fixed, considering the measures to limit the spread of the disease during the pandemic, namely lockdowns, masking, informing the population, testing rates, and vaccination. The residual plots showed accumulative deviation of the predictions and the actual measured data especially past the initial exponential part. That is to say, that even though the framework might appropriately be used to generate short term trends during stable environments, it is not adaptive in long-term simulation hence making use of real world dynamics that change quickly. The data have been criticized in that the recovery cases are underestimated due to shortcomings in recovery rate assumption and the quality of real world recovery reporting, which is incomplete or delayed in most cases.
Conclusion
The paper has been able to integrate a deterministic SEIR (SusceptibleExposedInfectiousRecovered) model in simulating the spread of COVID-19 and the overall predictive validity of the model was assessed using the comparison with real epidemiological data. The simulation was both useful and practical in the realization of the dynamic feature of infectious disease transmission but showed little in the way of prospective generalization over time. A fundamental weakness was the fixed parameter value which inhibited the model to include the impact of the modified public health interventions, the behavior adaptation, and altering patterns of transmission over a period. The negative values of R 2 and the high Root Mean Squared Error (RMSE) numbers in the quantitatively measured performance measures show that dynamic parameterization and the use of more flexible models would be necessary to achieve better accuracy and responsiveness to complexities in the real world.
To improve on the calibration of the model, it is suggested that, in a future study, one should use data-based parameter optimization methods, e.g. non-linear curve fitting, or machine learning methods as an optimization tool. Also, incorporation of time-varying parameters would help to better capture the changes in policy and individual reactions to an outbreak over a time of an epidemic. The current SEIR framework could be extended further to reflect our understanding of epidemic dynamics in much more detail, thereby improving our ability to simulate dynamic disease trajectories, and better inform policy making with respect to them, via additional compartments, including SEIR-V (or SEIR-QD) or SEIR-C (including consideration of contributions made by herd immunity as its own compartment or factor).
References
- Aganovic, A., Cao, G., Kurnitski, J., &Wargocki, P. (2023). New dose–response model and SARS-CoV-2 quanta emission rates for calculating the long-range airborne infection risk. Building and Environment, 228, Article 109924. https://doi.org/10.1016/j.buildenv.2022.109924
- Arifin, S. M. N., Arifin, R. R., Pitts, D. D. A., Rahman, M. S., Nowreen, S., Madey, G. R., & Collins, F. H. (2015). Landscape epidemiology modeling using an agent-based model and a geographic information system. Land, 4(2), 378–412. https://doi.org/10.3390/land4020378
- Becker, A. D., Grantz, K. H., Hegde, S. T., Bérubé, S., Cummings, D. A. T., & Wesolowski, A. (2021). Development and dissemination of infectious disease dynamic transmission models during the COVID-19 pandemic: What can we learn from other pathogens and how can we move forward? The Lancet. Digital Health, 3(1), e41–e50. https://doi.org/10.1016/S2589-7500(20)30268-5
- Bruch, E., & Atwell, J. (2015). Agent-based models in empirical social research. Sociological Methods and Research, 44(2), 186–221. https://doi.org/10.1177/0049124113506405
- Calvetti, D., Hoover, A. P., Rose, J., &Somersalo, E. (2020). Metapopulation network models for understanding, predicting, and managing the coronavirus disease COVID-19. Frontiers in Physics, 8, 261. https://doi.org/10.3389/fphy.2020.00261
- Chakroun, M., Haddad, S., Shakir, W. M. R., Sayah, J., Kallab, C., &Falou, S. (2024). Recent Monte Carlo-based simulations for modeling disease spread International Conference on Computer and Applications (ICCA). IEEE. https://doi.org/10.1109/ICCA62237.2024.10927993
- Das, S. Network-based modeling of infectious disease spillover. Diss. Kansas State University. (2024).
- Duan, W., Fan, Z., Zhang, P., Guo, G., & Qiu, X. (2015). Mathematical and computational approaches to epidemic modeling: A comprehensive review. Frontiers of Computer Science, 9(5), 806–826. https://doi.org/10.1007/s11704-014-3369-2
- Gallagher, S., & Baltimore, J. Comparing compartment and agent-based models Joint statistical meeting, Baltimore. 2017.
- Gao, S., Dai, X., Wang, L., Perra, N., & Wang, Z. (2022). Epidemic spreading in metapopulation networks coupled with awareness propagation. IEEE Transactions on Cybernetics, 1–13.
- Germano, D. P. J. et al. (2024). ‘A hybrid framework for compartmental models enabling simulation-based inference.’ arXiv preprint arXiv:2405.13239.
- Gomez, J., Prieto, J., Leon, E., & Rodríguez, A. (2021). INFEKTA—An agent-based model for transmission of infectious diseases: The COVID-19 case in Bogotá, Colombia. PLOS One, 16(2), Article e0245787. https://doi.org/10.1371/journal.pone.0245787
- Hunter, E., Mac Namee, B., & Kelleher, J. (2020). A hybrid agent-based and equation-based model for the spread of infectious diseases. Journal of Artificial Societies and Social Simulation, 23(4), 14. https://doi.org/10.18564/jasss.4421
- Lachi, A. (2024). Compartmental models and uncertainty quantification in epidemiology.
- Lancet Commission on Strengthening the Use of Epidemiological Modelling of Emerging and Pandemic Infectious Diseases. Electronic address: mark.jit@lshtm.ac.uk. (2024). How modelling can better support public health policy making: The Lancet Commission on strengthening the use of epidemiological modelling of emerging and pandemic infectious diseases. Lancet, 403(10429), 789–791. https://doi.org/10.1016/S0140-6736(23)02758-7
Statements & Declarations:
Peer-Review Method
This article underwent double-blind peer review by two external reviewers.
Competing Interests
The author/s declare no competing interests.
Funding
This research received no external funding.
Data Availability
Data are available from the corresponding author on reasonable request.
Licence
Simulation-Based Analysis of Disease Spread Using Compartmental Models © 2025 by Sonia Bansal & Sudesh Kumar is licensed under CC BY-NC-ND 4.0. Published by IJABS.
